DNA relaxes from UV excitation through several competing pathways.
In brief:
- We simulated the excited-state dynamics of alternating and nonalternating stacked guanine–cytosine tetramers.
- Both systems rapidly formed guanine-to-cytosine charge-transfer states after photoexcitation.
- The main relaxation channel involved interstrand proton transfer, but the electron and proton did not always move between the same bases.
DNA absorbs ultraviolet light. That is unavoidable. The relevant question is what happens next.
If the absorbed energy remains trapped for too long, chemical reactions can damage nucleobases, alter hydrogen-bonding patterns, or form photoproducts. Yet DNA is remarkably photostable. It has ways to quickly remove electronic excitation energy and return to the ground state before too much chemistry occurs.
For isolated nucleobases, the usual picture is relatively simple: the molecule distorts and reaches a conical intersection, a region where the excited and ground electronic states meet. But DNA is not a collection of isolated bases. Bases are paired by hydrogen bonds and stacked on top of one another. These contacts open other routes. Electrons may shift from one base to another. Protons may move across hydrogen bonds. Sometimes both happen in a coupled way.
The difficulty is that several mechanisms compete. Looking at one optimized structure is not enough. One needs dynamics.
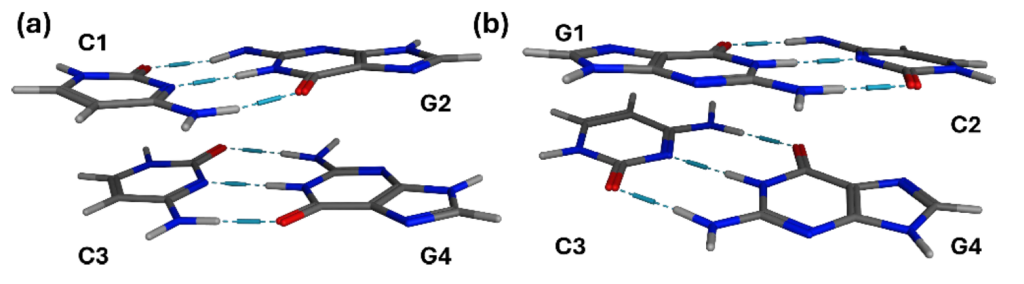
In a project developed by Juliana G. de Abrantes, we studied the photoexcited dynamics of minimal DNA models containing two stacked guanine–cytosine base pairs. We compared two arrangements: alternating and nonalternating GC tetramers. They are small enough to allow nonadiabatic dynamics, yet contain the two essential ingredients missing from monomers: hydrogen bonding and base stacking.
We used surface-hopping dynamics with TD-CAM-B3LYP and followed about 200 trajectories in total. The systems were initially excited around the lowest bright absorption band, near 4.9 eV. Very quickly, within the first tens of femtoseconds, the electronic excitation acquired charge-transfer character: electron density shifted primarily from guanine to cytosine.
Then the nuclei responded.
The dominant relaxation mechanism was proton transfer across the central guanine–cytosine hydrogen bond. This N–N proton transfer accounted for most of the relaxation events: 70% in the nonalternating tetramer and 82% in the alternating one. A second channel, involving N–O proton transfer from guanine to cytosine, was also important. The trajectories returned toward the ground state with characteristic times of about 64 fs for the nonalternating system and 141 fs for the alternating one.
The important point is not only that charge transfer precedes proton transfer. It is that the two motions are correlated without being locked together. In some trajectories, the electron and proton move between the same bases, resembling excited-state hydrogen transfer. In others, the electron follows a different route, especially in the alternating sequence, where π-stacking can support intrastrand charge transfer. Figure 4 of the paper (shown below) is useful here: it condenses a messy ensemble of trajectories into a small set of electron- and proton-transfer patterns.
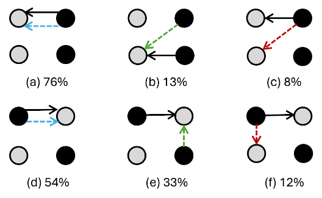
Scheme of main proton transfers, represented by solid arrows, and corresponding electron transfers, represented by dashed arrows, for nonalternating (a, b, and c) and alternating (d, e, and f) (GC)2 tetramers. Guanine is represented as a black circle and cytosine as a gray one. Under each scheme, a number denotes the percentage of the respective transfer pattern.
This result argues against a single-mechanism picture of DNA photostability. The same overall sequence—photoexcitation, charge separation, proton motion, relaxation—can unfold through several microscopic routes. The system is not confused. It is flexible.
There are limits. These are gas-phase tetramer models, with the backbone constrained. The mechanism is obtained within the present TDDFT-based surface-hopping framework, and solvent or full DNA environment effects may alter the pathway balance. Still, the work gives a dynamic view of how stacked GC bases can funnel UV excitation back to the ground state.
This paper should be cited for surface-hopping simulations of GC tetramers, for the coupled charge-transfer/proton-transfer mechanism in DNA photorelaxation, and for trajectory-based analysis of excited-state character using transition-density descriptors.
MB
Reference
[1] J. G. De Abrantes, J. M. Toldo, M. Barbatti, M. Sacchi, The Hidden Routes of DNA Photostability: Charge and Proton Transfer in Excited Cytosine–Guanine Tetramers, J. Phys. Chem. Lett. (2026). DOI: 10.1021/acs.jpclett.6c00376