Machine learning can reduce dynamics cost by a factor of ten.
In brief:
- Machine learning (ML) is used to predict energies and gradients during surface hopping simulations.
- ML delivers virtually exact results in one dimension.
- Applied to 33 dimensions, a training set with 10,000 points delivers results for 2 ps with reasonable accuracy, reducing costs by a factor of up to 10.
Machine learning (ML) has been one of the most exciting algorithmic developments of our time. It has enabled applications in every field of activity. It couldn’t be different in theoretical chemistry.
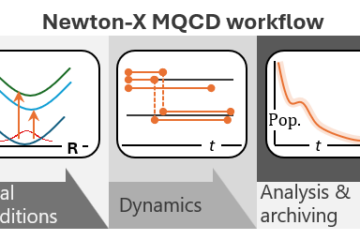
In a project led by Pavlo Dral, we built an interface between MLatom and Newton-X to evaluate the use of ML to predict energies, gradients, and couplings, the quantities needed to propagate decoherence-corrected fewest switches surface hopping (DC-FSSH).
The rationale is that if the number of quantum-mechanical points needed to train the machine is significantly smaller than the total number of points spanned by dynamics, then ML could reduce the heavy computational cost of dynamics propagation.
Based on the Kernel Ridge Regression (KRR) algorithm, we show that ML can in principle be used to accurately reproduce nonadiabatic excited-state dynamics based on DC-FSSH approach. The problem, of course, is that to achieve high accuracy in the prediction of the quantum mechanical results for multi-dimensional systems, we may need unaffordably large training sets. That’s the infamous curse of dimensionality.
The question is, is there some acceptable accuracy level we can reach with ML that still significantly reduces the simulation time of a realistic, high-dimensional system?
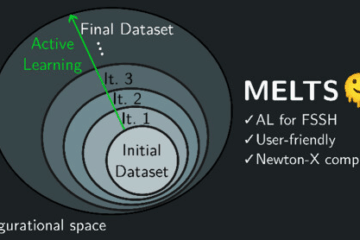
Our approach is based on creating approximate machine learning potentials for each adiabatic state on a relatively small number of training points, something like 1,000 to 10,000 points. Just for comparison, one single trajectory requires 10,000 points, and we may need 100 to 1,000 trajectories.
In our basic algorithm, ML is used only to predict energies and gradients in the regions with low hopping probability. The problem is that nonadiabatic couplings are extremely narrow functions, and their ML prediction would require too large training sets.
We have investigated the feasibility of this approach by using adiabatic spin-boson Hamiltonian (A-SBH) models of various dimensions as reference methods.
We learned that for 33-D, ML with 10,000 training points can deliver results for 2 ps dynamics with acceptable accuracy, reducing the number of QM points by a factor of six to eight, but that could be pushed to ten by controlling the thresholds.
These results are reported in Ref. [1].
MB
Reference
[1] P. O. Dral, M. Barbatti, and W. Thiel, Nonadiabatic Excited-State Dynamics with Machine Learning, J. Phys. Chem. Lett., 9, 5660 (2018). DOI: 10.1021/acs.jpclett.8b02469
- Pavlo has also written a post on this paper. You can see it here.