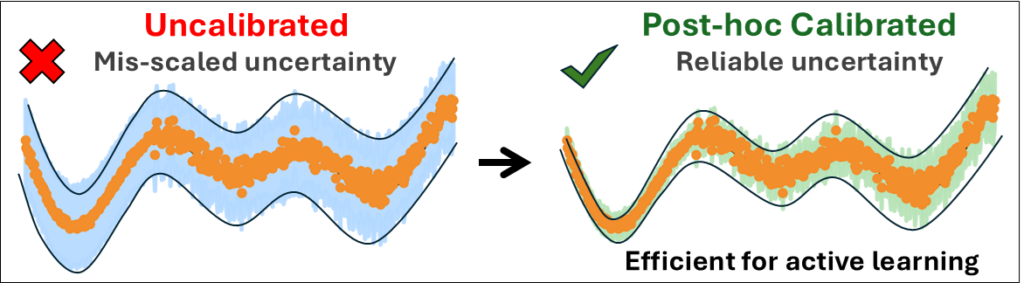
Post hoc calibration turns uncertainty estimates from decorative numbers into useful decision signals for chemistry ML.
In brief:
-
We benchmarked two popular uncertainty quantification (UQ) paradigms—Deep Evidential Regression and deep ensembles—on QM9 and WS22.
-
Raw uncertainties were systematically miscalibrated, but simple post hoc calibration methods fixed this without retraining the underlying models.
-
Calibration can save expensive reference calculations: in an active-learning-like test, we avoid thousands of redundant labels and cut selected training points by >20%.
Machine-learning models are now routine in quantum chemistry, from property prediction to building fast surrogate potentials. But there is a stubborn practical problem: when a model tells you it is “uncertain,” does that number actually correspond to being wrong? If not, you can waste serious compute time—especially when uncertainty is used to steer active learning, i.e., deciding which geometries deserve new quantum chemical calculations.
In a project led by Bidhan Chandra Garain, we tackled this head-on by focusing on calibration: adjusting predicted uncertainties so that (statistically) a stated confidence level matches observed errors. Put differently, calibration is the step that turns uncertainty into a quantity you can trust for decisions, not just plot in a figure.
We compared two widely used UQ paradigms:
The first is Deep Evidential Regression (DER), here implemented by modifying an E(n)-equivariant graph neural network (EGNN) to output an evidential distribution (Normal–Inverse–Gamma), which in principle separates epistemic (model) and aleatoric (data noise) contributions.
The second is the workhorse approach: deep ensembles, where multiple independently trained networks provide a predictive variance.
We then asked two questions.
First: Are the raw uncertainties calibrated? No. On both benchmarks, the native uncertainty outputs were systematically miscalibrated. DER in a way that blurred noise vs model uncertainty, ensembles in a way that could be “sharp” yet underconfident.
Second: Can we fix this cheaply, after training? Yes. We applied three post hoc calibration strategies spanning non-parametric and parametric families: isotonic regression, standard scaling, and GP-Normal.
These procedures adjust the uncertainty scale/mapping without changing the model’s mean prediction or retraining the network—so they are easy to slot into existing pipelines.
What changes once uncertainties are calibrated? On QM9, calibration made DER especially effective for confidence-based filtering: using a simple “high-confidence” threshold, calibrated DER selected thousands of predictions where low uncertainty genuinely corresponded to low error, whereas ensembles became overly conservative under the same rule.
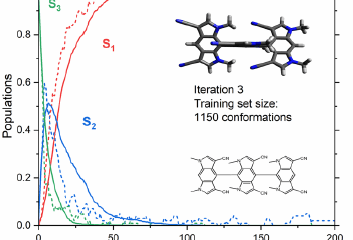
On WS22 (acrolein)—a more out-of-distribution setting built to stress-test generalization—calibrated ensemble uncertainty became materially more reliable and, crucially, more efficient for data selection. In a single-iteration active-learning-like experiment, isotonic-regression-calibrated selection achieved similar accuracy while using 4,546 fewer training points than an uncalibrated strategy, corresponding to >20% fewer redundant ab initio evaluations in that selection pool.
Bottom line: in molecular ML, uncertainty without calibration is often a misleading instrument. Calibration is the step that makes UQ operational—useful for triage, sampling, and compute-aware exploration of chemical space.
MB
Reference
[1] B. C. Garain, M. Pinheiro Jr., M. d. O. Bispo, M. Barbatti, Uncertainty Calibration in Molecular Machine Learning: Comparing Evidential and Ensemble Approaches, Chem. – Eur. J. e03299 (2026). DOI: 10.1002/chem.202503299